This article was automatically translated from the original Turkish version.
+1 More
Ribonucleic acid (RNA) is a high-molecular-weight biopolymer composed of ribonucleotides linked by phosphodiester bonds. Each nucleotide contains a nitrogenous base—adenine (A), guanine (G), cytosine (C), or uracil (U)—attached to a ribose sugar. RNA plays a central role in gene expression and cellular regulation, functioning as messenger, structural, regulatory molecules, and in some viruses as the genetic material in place of DNA.
All RNA molecules arise as transcripts of DNA sequences; however, not every transcript assumes a functional role. Therefore, a clear distinction must be made between functional RNAs and nonfunctional transcriptional byproducts to enable meaningful classification of RNA types.
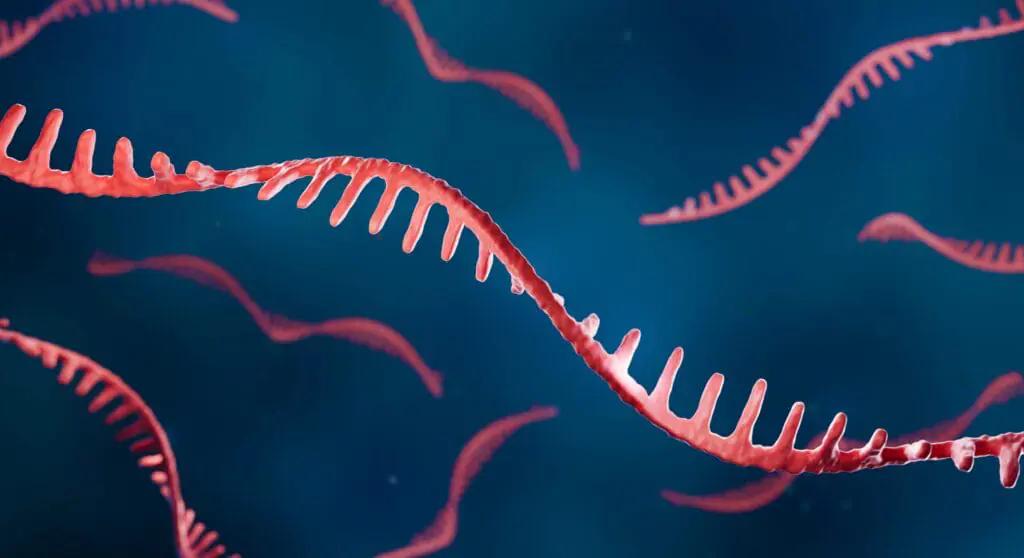
RNA Structure( yourgenome)
More than 60 years since the discovery of messenger RNA in 1961 and the development of the first FDA-approved mRNA vaccine in 2021, RNA research has undergone remarkable progress and transformation globally, particularly in the field of RNA therapeutics. Key discoveries such as RNA splicing (1977), self-splicing ribozymes (1982), and the first microRNA have paved the way for clinical applications. Antisense oligonucleotides (ASOs) reduce gene expression by specifically binding to target mRNAs, while aptamers behave like antibodies by binding to protein targets with high specificity and affinity. One of the greatest advantages of RNA-based drugs is their ability to target molecules previously considered “undruggable,” especially noncoding RNAs. Despite challenges such as instability, immune activation, and low cellular uptake, advances in delivery technologies are overcoming these limitations. Overall, RNA therapeutics are emerging as promising global tools for treating neurological, cardiovascular, genetic, cancer, and infectious diseases, offering faster, more cost-effective, and adaptable alternatives compared to traditional drug development approaches.
In Türkiye, significant research has been conducted in recent years in the fields of RNA-based detection and transcriptomic analysis. A notable example is the work led by Dr. Ersin Emre Ören from TOBB University of Economics and Technology in collaboration with U.S. institutions, in which atomic-scale electrodes were used to detect E. coli genetic material at the single-molecule level. This method enables the identification of single-nucleotide variations in DNA/RNA at attomolar concentrations, offering a novel approach for high-sensitivity genetic diagnostics. Additionally, a bibliometric mapping of sheep RNA sequencing and gene expression studies between 2011 and 2023 reveals limited publications from Türkiye, highlighting a research gap and future potential. Turkish researchers are encouraged to investigate breed-specific genetic variants, antiparasitic drug resistance, and climate-related animal traits, aligning with global scientific trends while addressing local agricultural and biomedical needs.
RNA is typically single-stranded but can form secondary and tertiary structures through intramolecular base pairing with complementary regions. These structures—such as bulges, loops, and helices—are stabilized by chemical modifications like methylation. Structural stability is critical for RNA function; unstable RNAs are typically nonfunctional and rapidly degraded by cellular quality control mechanisms.
Moreover, RNAs frequently associate with proteins to form ribonucleoprotein complexes (RNPs). In some cases, the RNA component itself exhibits catalytic activity, thereby assuming functions traditionally attributed only to proteins.
Methylation and secondary structure are two critical features that significantly contribute to RNA stability. The N7-guanosine (m⁷G) methylation at the 5′ end of mRNA forms the m⁷G cap, which protects the RNA from 5′-exonucleases and enhances overall stability. This cap also facilitates efficient translation and nuclear export. Meanwhile, RNA secondary structure—comprising hairpins, loops, and pseudoknots formed by internal base pairing—reduces accessibility to nucleases, thereby providing structural stability. Although mRNAs are largely linear, approximately 10–15% of nucleotides in eukaryotes participate in these local secondary structures; in viral RNAs, this proportion can exceed 30–40%. These structural elements protect RNA from enzymatic degradation and contribute to its functional integrity and lifespan by regulating interactions with cellular proteins.
Researchers in Türkiye have applied RNA sequencing technology to study T-cell acute lymphoblastic leukemia (T-ALL), a disease for which reliable prognostic genetic markers and a clear molecular classification are currently lacking. In a study utilizing Jurkat and Molt-4 cell lines, along with publicly available datasets (GSE48173) from healthy thymocyte subpopulations and T-ALL patients, a bioinformatics workflow was developed to analyze complex transcriptomic data. Findings revealed tissue-specific alternative splicing events, differences in gene expression, and significant genetic variations. This study reflects Türkiye’s advancing genomic research capacity through computational approaches that contribute to molecular diagnosis and therapeutic strategies.
To manage the increasing complexity observed in transcriptomic studies, a high-level classification system has been proposed. This model defines three main RNA superfamilies based on biological function:
These are RNA molecules with no defined or demonstrated function. This group includes background transcription products, degradation intermediates, splicing byproducts (such as introns), and nonfunctional readthrough transcripts. Although chemically RNA, they are not considered functional RNAs in a biological context and are termed “transcripts.”
RNA degradation plays a critical role in post-transcriptional gene regulation. As Houseley and Tollervey noted, early comparisons between RNA synthesis and steady-state levels revealed that cells produce more RNA than they retain, demonstrating the existence of active RNA degradation systems. MicroRNAs (miRNAs) also make significant contributions to gene expression regulation. According to Friedman and colleagues, over 45,000 conserved miRNA target sites exist in human 3′UTRs, and more than 60% of human protein-coding genes have been under evolutionary pressure to preserve miRNA binding. These findings establish RNA stability and miRNA-mediated repression as fundamental layers of gene expression regulation.
These RNAs assume structural, regulatory, catalytic, or carrier roles within the cell. Examples include transfer RNA (tRNA), ribosomal RNA (rRNA), small nuclear RNA (snRNA), small nucleolar RNA (snoRNA), microRNA (miRNA), and PIWI-interacting RNA (piRNA). Their biological activities, regardless of size or origin, define them as functional RNAs.
These are characterized by their ability to be translated into polypeptides. They contain open reading frames (ORFs) and untranslated regions (UTRs) that may include regulatory elements. They serve as templates for protein synthesis on ribosomes and stand out through both coding and regulatory functions.
In Türkiye, under the coordination of TÜSEB, significant advances have been made in transcriptomic and genomic research through the National Genome and Bioinformatics Project. As part of this initiative, the Türkiye Cancer Institute has launched a pioneering study titled “Personalized Medicine Approach in Bladder Cancer Treatment: Genomic/Transcriptomic Analyses and Tumor Modeling.” This project aims to integrate comprehensive genomic and transcriptomic data from bladder cancer patients with clinical and drug resistance information to create a functional genomic resource. Using Illumina platforms for high-depth whole-genome sequencing (WGS) and whole-transcriptome sequencing (WTS), the study also develops 3D tumor organoid cultures to evaluate therapeutic responses. This effort represents a foundational step in personalized oncology in Türkiye, with plans to expand to other cancers such as breast cancer, aiming to advance innovative treatment strategies and strengthen international scientific collaboration.
Boundaries between RNA classes are often fluid. For instance, some mRNAs partially overlap with Class I RNAs by containing noncoding regulatory elements within their UTRs. Conversely, some Class I RNAs occasionally contain short ORFs capable of translation, rendering them bifunctional RNAs.
Hybrid molecules also exist. For example, bacterial tmRNA combines tRNA and mRNA functions. Circular RNAs (circRNAs), formed when 5′ and 3′ ends are covalently linked into a circle, can act as miRNA sponges or serve as translation templates. However, the majority of circRNAs are produced through aberrant splicing, are nonfunctional, and are classified as Class 0 transcripts.
The naturally expressed circular RNA ciRS-7 acts as a potent sponge for miR-7, containing over 70 conserved binding sites; it is resistant to miRNA-mediated degradation and reduces miR-7 activity. Co-expression of ciRS-7 and miR-7 in the mouse brain indicates a functional interaction in neuronal tissues. Additionally, the testis-specific circRNA Sry behaves as a sponge for miR-138, suggesting that circRNA-mediated miRNA regulation may be a widespread endogenous mechanism.
The ssrA gene encodes tmRNA and is found in all sequenced eubacterial genomes and some plastid chromosomes. tmRNA molecules are typically 325–400 nucleotides long, but shorter forms exist in plastids and cyanobacteria due to reductive evolution. Structural variants are also observed in some α-proteobacteria and cyanobacteria, where tmRNA consists of two noncovalent RNA chains. tmRNA is transcribed as a precursor and processed into its mature form by RNases. The 5′ end is generated by RNase P; 3′ end maturation, as studied in E. coli, involves RNase E and exonuclease activity.
In a transcriptome analysis of hypothalamic and pituitary tissues from chickens with high and low egg production, 4,644 differentially expressed genes were identified. Chickens with low egg production showed upregulation of genes in the hypothalamic-pituitary-thyroid (HPT) axis, while high-producing chickens exhibited increased beta-estradiol activity. Thyroid hormone more strongly repressed gonadotropin mRNA in high-producing chickens, whereas estradiol stimulated gonadotropin mRNA in low-producing chickens but repressed it in high-producing ones. These findings highlight distinct hormonal regulation of the HPG axis in relation to egg production. Such results are valuable for advancing poultry breeding and reproductive research in Türkiye.
At the time of diagnosis, tumor stage significantly impacts recurrence, progression, and mortality in cancer patients. However, for many cancer types, effective early detection methods or screening tests are lacking, making microRNA-based diagnostics a promising field. MicroRNAs possess suitable characteristics as biomarkers, including cancer- and tissue-specific expression profiles and detectability in body fluids. Moreover, their chemical stability in fresh or formalin-fixed tissues and body fluids makes them more suitable than longer mRNAs or long noncoding RNAs. Since microRNAs are secreted by both healthy and cancerous cells, research has intensified to identify specific expression signatures in blood, urine, or fecal samples as potential diagnostic markers.
Cancer cells (Ministry of Health)
tRNAs and tRNA-derived fragments (tRFs) are also associated with diseases. tRNAs can inhibit apoptosis by interacting with caspase proteins, a mechanism linked to cancer cell survival. Repetitive RNA sequences that sequester RNA-binding proteins (RBPs), rendering them nonfunctional, are observed in neurological disorders such as amyotrophic lateral sclerosis (ALS) and myotonic dystrophy. Mutations and misregulation of RBPs contribute to numerous human diseases.
Advances in RNA sequencing and transcriptomic profiling have enabled the discovery of tumor-specific RNA biomarkers such as MALAT1, which is associated with cancer metastasis and cell proliferation.
MALAT1 is a conserved and abundant long noncoding RNA involved in gene regulation. Although no developmental defects are observed in mice lacking the MALAT1 gene, this RNA has been linked to metastasis in lung and various other cancers. The role of MALAT1 varies depending on cancer type and experimental model, and different inactivation methods can produce distinct effects. Multiple mechanisms have been proposed to explain MALAT1’s function in cancer progression.
Brady, K., H. C. Liu, J. A. Hicks, et al. “Transcriptome Analysis of the Hypothalamus and Pituitary of Turkey Hens with Low and High Egg Production.” *BMC Genomics* 21 (2020): 647. https://doi.org/10.1186/s12864-020-07075-y.
Brighton, Katie, and Eliza Small. *Addressing RNA Research Challenges*. Technology Networks, October 18, 2022. Accessed July 19, 2025. https://www.technologynetworks.com/biopharma/blog/addressing-rna-research-challenges-366380.
Brosius, Jürgen, and Carsten A. Raabe. “What Is an RNA? A Top Layer for RNA Classification.” *RNA Biology* 13, no. 2 (2016): 140–144. https://doi.org/10.1080/15476286.2015.1128064.
Duong, Hong-Quan, Minh-Cong Hoang, Thi-Hue Nguyen, Van-Lang Ngo, and Van-Thu Le. “Chapter Six - RNA Therapeutics History and Future Perspectives.” *Progress in Molecular Biology and Translational Science*, 2024. https://doi.org/10.1016/bs.pmbts.2024.01.004.
Friedman, R. C., K. K. Farh, C. B. Burge, and D. P. Bartel. “Most Mammalian mRNAs Are Conserved Targets of microRNAs.” *Genome Research* 19, no. 1 (January 2009): 92–105. https://doi.org/10.1101/gr.082701.108.
Hansen, T. B., T. I. Jensen, B. H. Clausen, J. B. Bramsen, B. Finsen, C. K. Damgaard, and J. Kjems. “Natural RNA Circles Function as Efficient microRNA Sponges.” *Nature* 495, no. 7441 (March 21, 2013): 384–388. https://doi.org/10.1038/nature11993.
Houseley, J., and D. Tollervey. “The Many Pathways of RNA Degradation.” *Cell* 136, no. 4 (February 20, 2009): 763–776. https://doi.org/10.1016/j.cell.2009.01.019.
Janssen, Brian D., and Christopher S. Hayes. “The tmRNA Ribosome Rescue System.” *Advances in Protein Chemistry and Structural Biology* 86 (2012): 151–191. https://doi.org/10.1016/B978-0-12-386497-0.00005-0.
Kornienko, Igor V., Olga Yu Aramova, Anna A. Tishchenko, Dmitriy V. Rudoy, and Michael Leonidas Chikindas. “RNA Stability: A Review of the Role of Structural Features and Environmental Conditions.” *Molecules* 29, no. 24 (December 18, 2024): 5978. https://doi.org/10.3390/molecules29245978.
Pichler, M., and G. A. Calin. “MicroRNAs in Cancer: From Developmental Genes in Worms to Their Clinical Application in Patients.” *British Journal of Cancer* 113 (2015): 569–573. https://doi.org/10.1038/bjc.2015.245.
Sun, Y., and L. Ma. “New Insights into Long Non-Coding RNA MALAT1 in Cancer and Metastasis.” Cancers (Basel) 11, no. 2 (February 13, 2019): 216. https://doi.org/10.3390/cancers11020216.
Turgutlu No. 6 Family Health Center. "Kanser Nedir?" *Genel Sağlık Bilgileri*. Accessed July 19, 2025. http://www.turgutlu6noluasm.com/index.php/saglik-bilgi/genel-saglik-bilgileri/item/258-kanser-nedir.
Türkiye Kanser Enstitüsü (TÜSEB). "TÜSEB Kanser Tedavisinde Kişisel Tıp Yaklaşımıyla Yeni Bir Dönem Başlatıyor." Türkiye Kanser Enstitüsü, September 10, 2024. Accessed July 19, 2025. https://tke.tuseb.gov.tr/haberler/tuseb-kanser-tedavisinde-kisisel-tip-yaklasimiyla-yeni-bir-donem-baslatiyor-20240909.
Wellcome Sanger Institute. “What is RNA?” Your Genome. Accessed July 19, 2025. https://www.yourgenome.org/theme/what-is-rna/.
No Discussion Added Yet
Start discussion for "RNA (Ribonucleic Acid)" article
Molecular Structure
Classification and Functional Hierarchy
Class 0 – Transcripts
Class I – Functional RNAs
Class II – Messenger RNAs (mRNAs)
Structural and Functional Diversity
RNA in Human Diseases